HQ Team
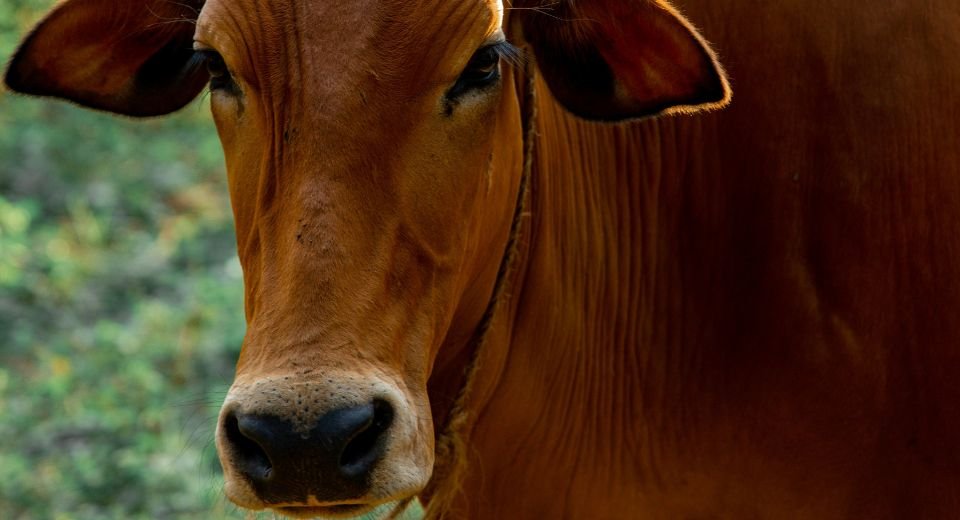
April 3, 2024: The deaths of about 100,000 cattle in India, which started two years ago, have been tracked to a viral infection, or lumpy skin disease outbreak, by scientists at the Indian Institute of Science, who said it was the second major outbreak in the South Asian nation.
“It was a calamity in some ways … a national emergency,” said Utpal Tatu, Professor in the Department of Biochemistry, Indian Institute of Science, in a statement. “The outbreak has severely affected India’s agricultural sector, leading to staggering economic losses.”
The disease caused by a virus — Lumpy Skin Disease Virus (LSDV) — is transmitted by insects such as flies and mosquitoes. It causes fever and skin nodules and can be fatal for cattle.
The virus was first found in Zambia in 1931 and remained confined to the Sub-African region until 1989, after which it started spreading to the Middle East, Russia and other southeast European nations, before spreading to South Asia.
There have been two major outbreaks of this disease in India, the first in 2019 and a more severe outbreak in May 2022, infecting more than two million cows, according to the IISc statement.
Veterinary institutes
The scientists collected skin nodules, blood and nasal swabs from infected cattle in various states, including Gujarat, Maharashtra, Rajasthan and Karnataka, in collaboration with veterinary institutes.
They performed advanced whole-genome sequencing of DNA extracted from 22 samples.
Genomic analysis revealed two distinct virus variants circulating in India, — one with a low number of genetic variations and another with a high number of genetic variations.
The sequence with fewer variations was genetically similar to the 2019 Ranchi (in the northern state of Jharkhand) and 2020 Hyderabad (southern India) strains that were sequenced previously.
The samples with high variations, however, turned out to be similar to LSDV strains from an outbreak in Russia in 2015.
‘Lack of LSDV genome sequencing’
“The biggest challenge was the lack of an established LSDV genome sequencing and analysis pipeline. We had to adapt techniques from COVID-19 research,” said Ankeet Kumar, PhD student at IISc and co-lead author. “Data was also limited, so we compiled all available global LSDV genome sequences to make our analysis robust.”
The team found a large number of genetic variations of more than 1,800. These include deletions and insertions in various genes, single-letter changes in DNA (called SNPs), and genetic variations in regions between genes.
They found a large number of genetic variations in viral genes critical for binding to host cells, evading immune response, and replicating efficiently. This likely enhanced the virus’s ability to cause disease.
“Cattle developed more severe symptoms in areas where we found highly diverse strains. This suggests that the genetic variations could elevate virulence,” Kumar said.
‘Surprising’
He said there are no previous reports of such highly varied LSDV strains in India. Viruses that have DNA as the genetic material – like LSDV – are generally more stable than RNA viruses. Therefore, finding so many genetic variations was quite “surprising, and could explain the severity of the disease.”
Professor Tatu said: “The genomic data will prove invaluable for vaccine development by revealing molecular hotspots and genetic variations to target.”
“This is a first for characterising the genomic landscape of LSDV during India’s outbreak on a national scale.”